| Record Information |
|---|
| Version | 1.0 |
|---|
| Created at | 2020-04-17 19:14:58 UTC |
|---|
| Updated at | 2020-12-07 19:11:41 UTC |
|---|
| CannabisDB ID | CDB005156 |
|---|
| Secondary Accession Numbers | Not Available |
|---|
| Cannabis Compound Identification |
|---|
| Common Name | (S)-3-Hydroxyisobutyryl-CoA |
|---|
| Description | (S)-3-Hydroxyisobutyryl-CoA, also known as 3-hydroxy-isobutyryl-coenzyme A or 3-hydroxy-2-methylpropanoyl-CoA, belongs to the class of organic compounds known as acyl coas. These are organic compounds containing a coenzyme A substructure linked to an acyl chain (S)-3-Hydroxyisobutyryl-CoA is a strong basic compound (based on its pKa). (S)-3-Hydroxyisobutyryl-CoA is expected to be in Cannabis as all living plants are known to produce and metabolize it. |
|---|
| Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| (S)-3-Hydroxyisobutyryl-coenzyme A | HMDB | | 3-Hydroxy-2-methylpropanoyl-CoA | HMDB | | 3-Hydroxy-2-methylpropanoyl-coenzyme A | HMDB | | 3-Hydroxy-2-methylpropionyl-CoA | HMDB | | 3-Hydroxy-2-methylpropionyl-coenzyme A | HMDB | | 3-Hydroxy-isobutyryl-CoA | HMDB | | 3-Hydroxy-isobutyryl-coenzyme A | HMDB | | 3-Hydroxyisobutyryl-CoA | HMDB | | 3-Hydroxyisobutyryl-coenzyme A | HMDB |
|
|---|
| Chemical Formula | C25H42N7O18P3S |
|---|
| Average Molecular Weight | 853.62 |
|---|
| Monoisotopic Molecular Weight | 853.152 |
|---|
| IUPAC Name | {[5-(6-amino-9H-purin-9-yl)-4-hydroxy-2-({[hydroxy({[hydroxy(3-hydroxy-3-{[2-({2-[(3-hydroxy-2-methylpropanoyl)sulfanyl]ethyl}carbamoyl)ethyl]carbamoyl}-2,2-dimethylpropoxy)phosphoryl]oxy})phosphoryl]oxy}methyl)oxolan-3-yl]oxy}phosphonic acid |
|---|
| Traditional Name | [5-(6-aminopurin-9-yl)-4-hydroxy-2-({[hydroxy([hydroxy(3-hydroxy-3-{[2-({2-[(3-hydroxy-2-methylpropanoyl)sulfanyl]ethyl}carbamoyl)ethyl]carbamoyl}-2,2-dimethylpropoxy)phosphoryl]oxy)phosphoryl]oxy}methyl)oxolan-3-yl]oxyphosphonic acid |
|---|
| CAS Registry Number | 319440-43-2 |
|---|
| SMILES | CC(CO)C(=O)SCCNC(=O)CCNC(=O)C(O)C(C)(C)COP(O)(=O)OP(O)(=O)OCC1OC(C(O)C1OP(O)(O)=O)N1C=NC2=C(N)N=CN=C12 |
|---|
| InChI Identifier | InChI=1S/C25H42N7O18P3S/c1-13(8-33)24(38)54-7-6-27-15(34)4-5-28-22(37)19(36)25(2,3)10-47-53(44,45)50-52(42,43)46-9-14-18(49-51(39,40)41)17(35)23(48-14)32-12-31-16-20(26)29-11-30-21(16)32/h11-14,17-19,23,33,35-36H,4-10H2,1-3H3,(H,27,34)(H,28,37)(H,42,43)(H,44,45)(H2,26,29,30)(H2,39,40,41) |
|---|
| InChI Key | WWEOGFZEFHPUAM-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as acyl coas. These are organic compounds containing a coenzyme A substructure linked to an acyl chain. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Fatty Acyls |
|---|
| Sub Class | Fatty acyl thioesters |
|---|
| Direct Parent | Acyl CoAs |
|---|
| Alternative Parents | |
|---|
| Substituents | - Coenzyme a or derivatives
- Purine ribonucleoside diphosphate
- Purine ribonucleoside bisphosphate
- Purine ribonucleoside 3',5'-bisphosphate
- Ribonucleoside 3'-phosphate
- Pentose-5-phosphate
- Pentose phosphate
- Beta amino acid or derivatives
- Glycosyl compound
- N-glycosyl compound
- 6-aminopurine
- Monosaccharide phosphate
- Organic pyrophosphate
- Imidazopyrimidine
- Purine
- Monoalkyl phosphate
- Aminopyrimidine
- Fatty amide
- Monosaccharide
- N-acyl-amine
- N-substituted imidazole
- Organic phosphoric acid derivative
- Imidolactam
- Phosphoric acid ester
- Alkyl phosphate
- Pyrimidine
- Tetrahydrofuran
- Azole
- Heteroaromatic compound
- Imidazole
- Carbothioic s-ester
- Secondary carboxylic acid amide
- Secondary alcohol
- Thiocarboxylic acid ester
- Carboxamide group
- Amino acid or derivatives
- Thiocarboxylic acid or derivatives
- Sulfenyl compound
- Carboxylic acid derivative
- Organoheterocyclic compound
- Oxacycle
- Azacycle
- Hydrocarbon derivative
- Organic nitrogen compound
- Organic oxide
- Organosulfur compound
- Organonitrogen compound
- Primary amine
- Organopnictogen compound
- Carbonyl group
- Organic oxygen compound
- Primary alcohol
- Organooxygen compound
- Alcohol
- Amine
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Ontology |
|---|
|
| Disposition | Route of exposure: Source: Biological location: |
|---|
| Role | Biological role: Industrial application: |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Properties | | Property | Value | Reference |
|---|
| Melting Point | Not Available | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | Not Available | Not Available | | logP | Not Available | Not Available |
|
|---|
| Predicted Properties | [] |
|---|
| Spectra |
|---|
| EI-MS/GC-MS | |
|---|
| MS/MS | | Type | Description | Splash Key | View |
|---|
| Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-000i-1912000120-c718f8aabcd5b3328193 | 2015-09-14 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-000i-1913000000-86eb8b4e6cdee60b559a | 2015-09-14 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-000i-2911000000-2d6f48b3506835c89ffd | 2015-09-14 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-00o9-9730140560-056ac34ed9b661751c57 | 2015-09-15 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-001i-3920110010-37a68eba22e3eecc9757 | 2015-09-15 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-057i-6900100000-e33bac5e5de19c7b719b | 2015-09-15 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0udi-0000000090-9147721d7b7a39a06164 | 2021-09-22 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0fri-1211103790-2282fa97e4f5351f2c0e | 2021-09-22 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0002-0129000000-625efeaf22c52212984a | 2021-09-22 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0udi-0000000090-ff327f166604b01cb3f3 | 2021-09-22 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0udr-9800102130-8d5a197de5930d76b37c | 2021-09-22 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-00p1-9103505710-c610ab0387d6ed968440 | 2021-09-22 | View Spectrum |
|
|---|
| NMR | | Type | Description | | View |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | | Spectrum |
|
|---|
| Pathways |
|---|
| Pathways | | Name | SMPDB/Pathwhiz | KEGG | | Valine, Leucine and Isoleucine Degradation |    | 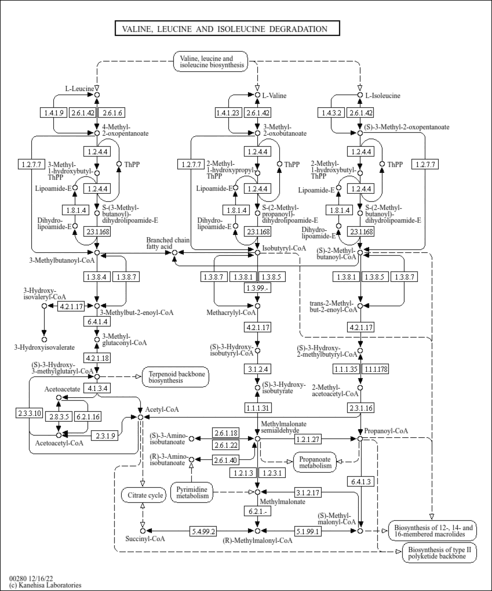 | | Beta-Ketothiolase Deficiency |  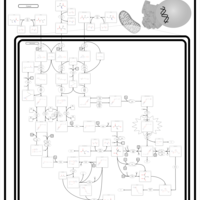 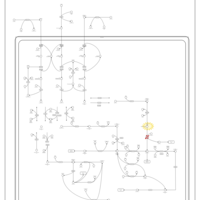 | Not Available | | 2-Methyl-3-Hydroxybutryl CoA Dehydrogenase Deficiency | 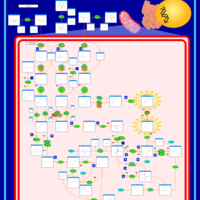 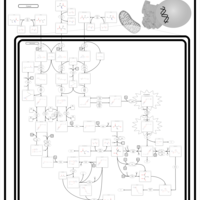 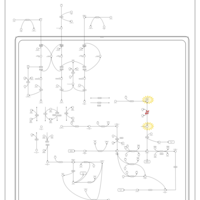 | Not Available | | Propionic Acidemia | 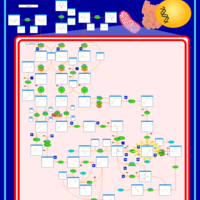 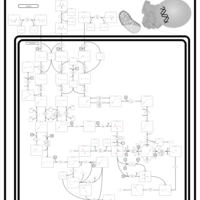 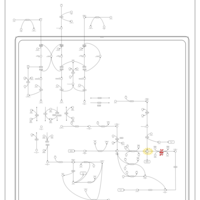 | Not Available | | 3-Hydroxy-3-Methylglutaryl-CoA Lyase Deficiency | 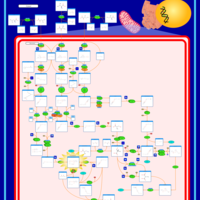 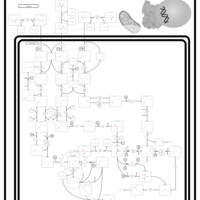 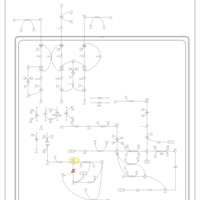 | Not Available |
|
|---|
| Protein Targets |
|---|
| Enzymes | |
| Enoyl-CoA hydratase, mitochondrial | ECHS1 | 10q26.2-q26.3 | P30084 | details | | 3-hydroxyisobutyryl-CoA hydrolase, mitochondrial | HIBCH | 2q32.2 | Q6NVY1 | details |
|
|---|
| Transporters | Not Available |
|---|
| Metal Bindings | Not Available |
|---|
| Receptors | Not Available |
|---|
| Transcriptional Factors | Not Available |
|---|
| Concentrations Data |
|---|
| Not Available |
|---|
| External Links |
|---|
| HMDB ID | HMDB0001052 |
|---|
| DrugBank ID | Not Available |
|---|
| Phenol Explorer Compound ID | Not Available |
|---|
| FoodDB ID | FDB022393 |
|---|
| KNApSAcK ID | Not Available |
|---|
| Chemspider ID | 86 |
|---|
| KEGG Compound ID | C06000 |
|---|
| BioCyc ID | Not Available |
|---|
| BiGG ID | 47176 |
|---|
| Wikipedia Link | Not Available |
|---|
| METLIN ID | 453 |
|---|
| PubChem Compound | 88 |
|---|
| PDB ID | Not Available |
|---|
| ChEBI ID | 28259 |
|---|
| References |
|---|
| General References | Not Available |
|---|
