| Identification |
|---|
| HMDB Protein ID
| CDBP05357 |
| Secondary Accession Numbers
| Not Available |
| Name
| Phosphatidate phosphatase LPIN1 |
| Description
| Not Available |
| Synonyms
|
- Lipin-1
|
| Gene Name
| LPIN1 |
| Protein Type
| Enzyme |
| Biological Properties |
|---|
| General Function
| Not Available |
| Specific Function
| Plays important roles in controlling the metabolism of fatty acids at differents levels. Acts as a magnesium-dependent phosphatidate phosphatase enzyme which catalyzes the conversion of phosphatidic acid to diacylglycerol during triglyceride, phosphatidylcholine and phosphatidylethanolamine biosynthesis in the reticulum endoplasmic membrane. Acts also as a nuclear transcriptional coactivator for PPARGC1A/PPARA to modulate lipid metabolism gene expression (By similarity). Is involved in adipocyte differentiation. May also be involved in mitochondrial fission by converting phosphatidic acid to diacylglycerol (By similarity).
|
| GO Classification
|
| Biological Process |
| fatty acid catabolic process |
| phosphatidylethanolamine biosynthetic process |
| phosphatidylcholine biosynthetic process |
| positive regulation of histone deacetylation |
| regulation of fat cell differentiation |
| ruffle organization |
| triglyceride mobilization |
| cellular response to insulin stimulus |
| negative regulation of transcription from RNA polymerase II promoter |
| positive regulation of transcription from RNA polymerase II promoter |
| actin cytoskeleton reorganization |
| triglyceride biosynthetic process |
| transcription, DNA-dependent |
| mitochondrial fission |
| organ regeneration |
| Cellular Component |
| transcription factor complex |
| endoplasmic reticulum membrane |
| cytosol |
| nuclear membrane |
| Molecular Function |
| phosphatidate phosphatase activity |
| transcription coactivator activity |
|
| Cellular Location
|
Not Available
|
| Pathways
|
| Name | SMPDB/Pathwhiz | KEGG | | Glycerophospholipid metabolism | Not Available | 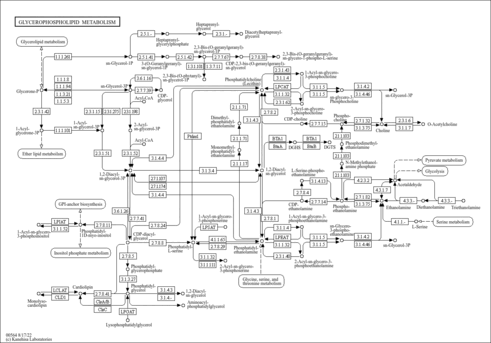 | | De Novo Triacylglycerol Biosynthesis |    | Not Available | | De Novo Triacylglycerol Biosynthesis TG(10:0/10:0/10:0) | 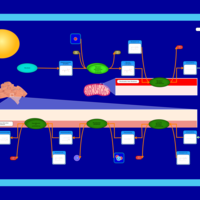 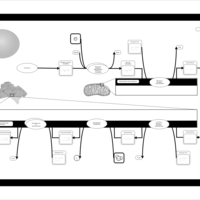 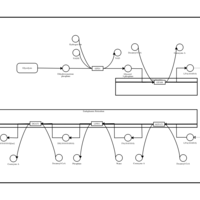 | Not Available | | De Novo Triacylglycerol Biosynthesis TG(16:0/16:0/16:0) | 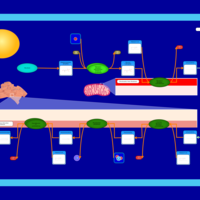 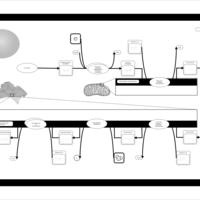 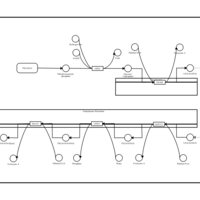 | Not Available | | De Novo Triacylglycerol Biosynthesis TG(16:0/16:0/18:0) | 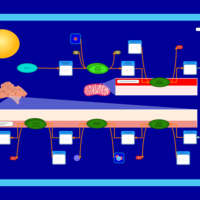 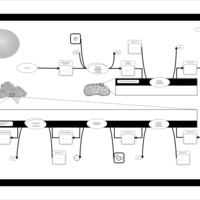  | Not Available |
|
| Gene Properties |
|---|
| Chromosome Location
| 2 |
| Locus
| 2p25.1 |
| SNPs
| Not Available |
| Gene Sequence
|
Not Available
|
| Protein Properties |
|---|
| Number of Residues
| Not Available |
| Molecular Weight
| 99366.085 |
| Theoretical pI
| 6.656 |
| Pfam Domain Function
|
|
| Signals
|
Not Available
|
|
Transmembrane Regions
|
Not Available
|
| Protein Sequence
|
>gi|387528011|ref|NP_001248356.1| phosphatidate phosphatase LPIN1 isoform 2 [Homo sapiens]
MSRVQTMNYVGQLAGQVFVTVKELYKGLNPATLSGCIDIIVIRQPNGNLQCSPFHVRFGK
MGVLRSREKV
|
| External Links |
|---|
| GenBank ID Protein
| Not Available |
| UniProtKB/Swiss-Prot ID
| Q14693 |
| UniProtKB/Swiss-Prot Entry Name
| Not Available |
| PDB IDs
|
Not Available |
| GenBank Gene ID
| Not Available |
| GeneCard ID
| Not Available |
| GenAtlas ID
| Not Available |
| HGNC ID
| HGNC:13345 |
| References |
|---|
| General References
| Not Available |
