| Identification |
|---|
| HMDB Protein ID
| CDBP05245 |
| Secondary Accession Numbers
| Not Available |
| Name
| Probable ATP-dependent RNA helicase DDX6 |
| Description
| Not Available |
| Synonyms
|
- ATP-dependent RNA helicase p54
- DEAD box protein 6
- Oncogene RCK
|
| Gene Name
| DDX6 |
| Protein Type
| Enzyme |
| Biological Properties |
|---|
| General Function
| Not Available |
| Specific Function
| In the process of mRNA degradation, may play a role in mRNA decapping.
|
| GO Classification
|
| Biological Process |
| exonucleolytic nuclear-transcribed mRNA catabolic process involved in deadenylation-dependent decay |
| gene expression |
| cytoplasmic mRNA processing body assembly |
| Cellular Component |
| cytosol |
| cytoplasmic mRNA processing body |
| RNA-induced silencing complex |
| cytoplasmic stress granule |
| Molecular Function |
| RNA helicase activity |
| ATP binding |
| ATP-dependent helicase activity |
| RNA binding |
|
| Cellular Location
|
Not Available
|
| Pathways
|
| Name | SMPDB/Pathwhiz | KEGG | | RNA degradation | Not Available | 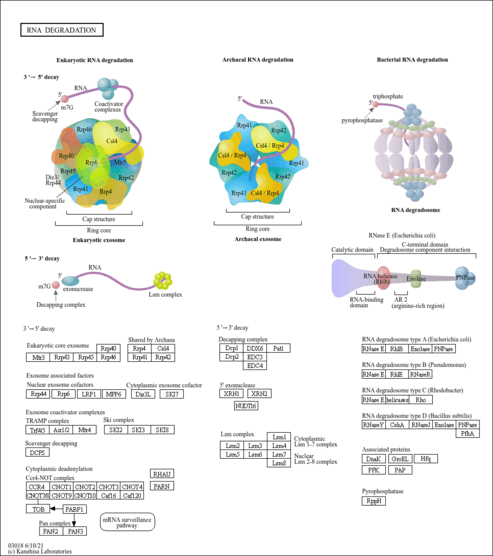 |
|
| Gene Properties |
|---|
| Chromosome Location
| 11 |
| Locus
| 11q23.3 |
| SNPs
| Not Available |
| Gene Sequence
|
Not Available
|
| Protein Properties |
|---|
| Number of Residues
| Not Available |
| Molecular Weight
| 54416.42 |
| Theoretical pI
| 8.666 |
| Pfam Domain Function
|
Not Available |
| Signals
|
Not Available
|
|
Transmembrane Regions
|
Not Available
|
| Protein Sequence
|
>gi|380692342|ref|NP_001244120.1| probable ATP-dependent RNA helicase DDX6 [Homo sapiens]
MSTARTENPVIMGLSSQNGQLRGPVKPTGGPGGGGTQTQQQMNQLKNTNTINNGTQQQAQ
SMTTTIKPGD
|
| External Links |
|---|
| GenBank ID Protein
| Not Available |
| UniProtKB/Swiss-Prot ID
| P26196 |
| UniProtKB/Swiss-Prot Entry Name
| Not Available |
| PDB IDs
|
|
| GenBank Gene ID
| Not Available |
| GeneCard ID
| Not Available |
| GenAtlas ID
| Not Available |
| HGNC ID
| HGNC:2747 |
| References |
|---|
| General References
| Not Available |
