| Identification |
|---|
| HMDB Protein ID
| CDBP05242 |
| Secondary Accession Numbers
| Not Available |
| Name
| Probable ATP-dependent RNA helicase DDX5 |
| Description
| Not Available |
| Synonyms
|
- DEAD box protein 5
- RNA helicase p68
|
| Gene Name
| DDX5 |
| Protein Type
| Enzyme |
| Biological Properties |
|---|
| General Function
| Not Available |
| Specific Function
| Involved in the alternative regulation of pre-mRNA splicing; its RNA helicase activity is necessary for increasing tau exon 10 inclusion and occurs in a RBM4-dependent manner. Binds to the tau pre-mRNA in the stem-loop region downstream of exon 10. The rate of ATP hydrolysis is highly stimulated by single-stranded RNA. Involved in transcriptional regulation; the function is independent of the RNA helicase activity. Transcriptional coactivator for estrogen receptor ESR1 and androgen receptor AR. Increases ESR1 AF-1 domain-mediated transactivation and ESR1 AF-1 and AF-2 domains transcriptional synergistic activity. Synergizes with DDX17 and SRA1 RNA to activate MYOD1 transcriptional activity and involved in skeletal muscle differentiation. Transcriptional coactivator for p53/TP53 and involved in p53/TP53 transcriptional response to DNA damage and p53/TP53-dependent apoptosis. Transcriptional coactivator for RUNX2 and involved in regulation of osteoblast differentiation. Acts as transcriptional repressor in a promoter-specicic manner; the function probbaly involves association with histone deacetylases, such as HDAC1.
|
| GO Classification
|
| Biological Process |
| negative regulation of transcription from RNA polymerase II promoter |
| positive regulation of transcription from RNA polymerase II promoter |
| transcription, DNA-dependent |
| positive regulation of intracellular estrogen receptor signaling pathway |
| regulation of skeletal muscle cell differentiation |
| in utero embryonic development |
| mRNA splicing, via spliceosome |
| cell growth |
| intrinsic apoptotic signaling pathway by p53 class mediator |
| positive regulation of DNA damage response, signal transduction by p53 class mediator |
| regulation of alternative mRNA splicing, via spliceosome |
| regulation of androgen receptor signaling pathway |
| regulation of osteoblast differentiation |
| regulation of viral genome replication |
| Cellular Component |
| nucleolus |
| catalytic step 2 spliceosome |
| Molecular Function |
| RNA helicase activity |
| ATP-dependent RNA helicase activity |
| ATP binding |
| estrogen receptor binding |
| transcription coactivator activity |
| androgen receptor binding |
| pre-mRNA binding |
|
| Cellular Location
|
Not Available
|
| Pathways
|
| Name | SMPDB/Pathwhiz | KEGG | | Spliceosome | Not Available | 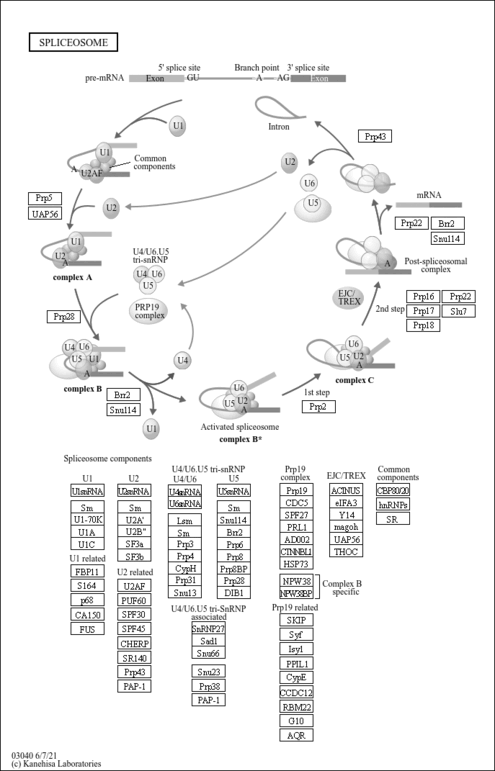 | | Transcriptional misregulation in cancer | Not Available | 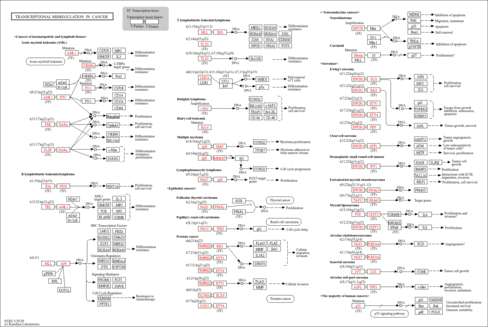 | | Proteoglycans in cancer | Not Available | 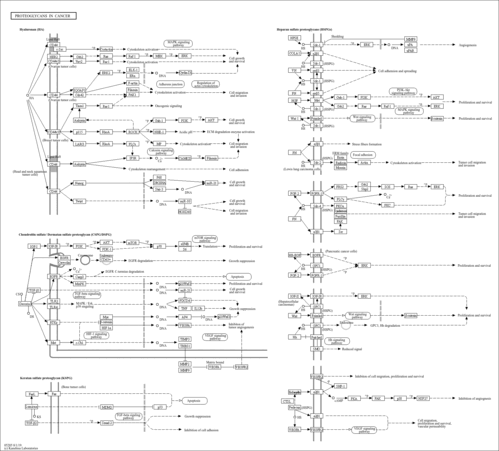 |
|
| Gene Properties |
|---|
| Chromosome Location
| 17 |
| Locus
| 17q21 |
| SNPs
| Not Available |
| Gene Sequence
|
Not Available
|
| Protein Properties |
|---|
| Number of Residues
| Not Available |
| Molecular Weight
| 69147.585 |
| Theoretical pI
| 8.924 |
| Pfam Domain Function
|
|
| Signals
|
Not Available
|
|
Transmembrane Regions
|
Not Available
|
| Protein Sequence
|
>gi|4758138|ref|NP_004387.1| probable ATP-dependent RNA helicase DDX5 [Homo sapiens]
MSGYSSDRDRGRDRGFGAPRFGGSRAGPLSGKKFGNPGEKLVKKKWNLDELPKFEKNFYQ
EHPDLARRTA
|
| External Links |
|---|
| GenBank ID Protein
| Not Available |
| UniProtKB/Swiss-Prot ID
| P17844 |
| UniProtKB/Swiss-Prot Entry Name
| Not Available |
| PDB IDs
|
|
| GenBank Gene ID
| Not Available |
| GeneCard ID
| Not Available |
| GenAtlas ID
| Not Available |
| HGNC ID
| HGNC:2746 |
| References |
|---|
| General References
| Not Available |


