| Identification |
|---|
| HMDB Protein ID
| CDBP05206 |
| Secondary Accession Numbers
| Not Available |
| Name
| m7GpppX diphosphatase |
| Description
| Not Available |
| Synonyms
|
- DCS-1
- Decapping scavenger enzyme
- Hint-related 7meGMP-directed hydrolase
- Histidine triad nucleotide-binding protein 5
- Histidine triad protein member 5
- Scavenger mRNA-decapping enzyme DcpS
- HINT-5
|
| Gene Name
| DCPS |
| Protein Type
| Enzyme |
| Biological Properties |
|---|
| General Function
| Not Available |
| Specific Function
| Decapping scavenger enzyme that catalyzes the cleavage of a residual cap structure following the degradation of mRNAs by the 3'->5' exosome-mediated mRNA decay pathway. Hydrolyzes cap analog structures like 7-methylguanosine nucleoside triphosphate (m7GpppG) with up to 10 nucleotide substrates (small capped oligoribonucleotides) and specifically releases 5'-phosphorylated RNA fragments and 7-methylguanosine monophosphate (m7GMP). Cleaves cap analog structures like tri-methyl guanosine nucleoside triphosphate (m3(2,2,7)GpppG) with very poor efficiency. Does not hydrolyze unmethylated cap analog (GpppG) and shows no decapping activity on intact m7GpppG-capped mRNA molecules longer than 25 nucleotides. Does not hydrolyze 7-methylguanosine diphosphate (m7GDP) to m7GMP (PubMed:22985415). May also play a role in the 5'->3 mRNA decay pathway; m7GDP, the downstream product released by the 5'->3' mRNA mediated decapping activity, may be also converted by DCPS to m7GMP (PubMed:14523240). Binds to m7GpppG and strongly to m7GDP. Plays a role in first intron splicing of pre-mRNAs. Inhibits activation-induced cell death.
|
| GO Classification
|
| Biological Process |
| exonucleolytic nuclear-transcribed mRNA catabolic process involved in deadenylation-dependent decay |
| cellular response to menadione |
| deadenylation-dependent decapping of nuclear-transcribed mRNA |
| mRNA cis splicing, via spliceosome |
| negative regulation of programmed cell death |
| Cellular Component |
| nucleus |
| cytosol |
| mitochondrion |
| Molecular Function |
| m7G(5')pppN diphosphatase activity |
| RNA 7-methylguanosine cap binding |
|
| Cellular Location
|
Not Available
|
| Pathways
|
| Name | SMPDB/Pathwhiz | KEGG | | RNA degradation | Not Available | 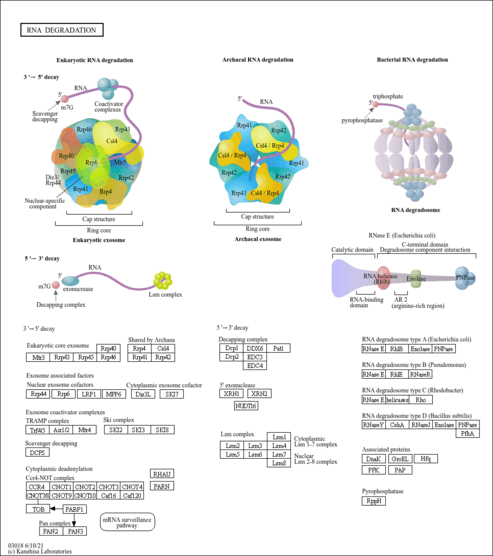 |
|
| Gene Properties |
|---|
| Chromosome Location
| 11 |
| Locus
| 11q24.2 |
| SNPs
| Not Available |
| Gene Sequence
|
Not Available
|
| Protein Properties |
|---|
| Number of Residues
| Not Available |
| Molecular Weight
| 38608.45 |
| Theoretical pI
| 6.377 |
| Pfam Domain Function
|
|
| Signals
|
Not Available
|
|
Transmembrane Regions
|
Not Available
|
| Protein Sequence
|
>gi|7661734|ref|NP_054745.1| m7GpppX diphosphatase [Homo sapiens]
MADAAPQLGKRKRELDVEEAHAASTEEKEAGVGNGTCAPVRLPFSGFRLQKVLRESARDK
IIFLHGKVNE
|
| External Links |
|---|
| GenBank ID Protein
| Not Available |
| UniProtKB/Swiss-Prot ID
| Q96C86 |
| UniProtKB/Swiss-Prot Entry Name
| Not Available |
| PDB IDs
|
|
| GenBank Gene ID
| Not Available |
| GeneCard ID
| Not Available |
| GenAtlas ID
| Not Available |
| HGNC ID
| HGNC:29812 |
| References |
|---|
| General References
| Not Available |
