| Identification |
|---|
| HMDB Protein ID
| CDBP00856 |
| Secondary Accession Numbers
| Not Available |
| Name
| Poly [ADP-ribose] polymerase 1 |
| Description
| Not Available |
| Synonyms
|
- ADPRT 1
- NAD(+) ADP-ribosyltransferase 1
- PARP-1
- Poly[ADP-ribose] synthase 1
- ADP-ribosyltransferase diphtheria toxin-like 1
- ARTD1
|
| Gene Name
| PARP1 |
| Protein Type
| Enzyme |
| Biological Properties |
|---|
| General Function
| Involved in DNA binding |
| Specific Function
| Involved in the base excision repair (BER) pathway, by catalyzing the poly(ADP-ribosyl)ation of a limited number of acceptor proteins involved in chromatin architecture and in DNA metabolism. This modification follows DNA damages and appears as an obligatory step in a detection/signaling pathway leading to the reparation of DNA strand breaks. Mediates the poly(ADP-ribosyl)ation of APLF and CHFR. Positively regulates the transcription of MTUS1 and negatively regulates the transcription of MTUS2/TIP150. With EEF1A1 and TXK, forms a complex that acts as a T-helper 1 (Th1) cell-specific transcription factor and binds the promoter of IFN-gamma to directly regulate its transcription, and is thus involved importantly in Th1 cytokine production.
|
| GO Classification
|
| Biological Process |
| transforming growth factor beta receptor signaling pathway |
| transcription initiation from RNA polymerase II promoter |
| protein autoprocessing |
| DNA repair |
| cellular response to insulin stimulus |
| telomere maintenance |
| negative regulation of transcription from RNA polymerase II promoter |
| base-excision repair |
| protein poly-ADP-ribosylation |
| DNA damage response, detection of DNA damage |
| regulation of growth rate |
| Cellular Component |
| nucleolus |
| nuclear envelope |
| transcription factor complex |
| Component |
| cell part |
| organelle |
| membrane-bounded organelle |
| intracellular membrane-bounded organelle |
| nucleus |
| intracellular |
| Function |
| catalytic activity |
| transition metal ion binding |
| zinc ion binding |
| transferase activity |
| nad+ adp-ribosyltransferase activity |
| nad or nadh binding |
| transferase activity, transferring pentosyl groups |
| nucleic acid binding |
| dna binding |
| transferase activity, transferring glycosyl groups |
| ion binding |
| cation binding |
| metal ion binding |
| binding |
| nucleotide binding |
| Molecular Function |
| NAD binding |
| metal ion binding |
| NAD+ ADP-ribosyltransferase activity |
| zinc ion binding |
| DNA binding |
| Process |
| macromolecule metabolic process |
| protein amino acid adp-ribosylation |
| post-translational protein modification |
| macromolecule modification |
| protein modification process |
| metabolic process |
|
| Cellular Location
|
- Nucleus
|
| Pathways
|
| Name | SMPDB/Pathwhiz | KEGG | | Base excision repair | Not Available | 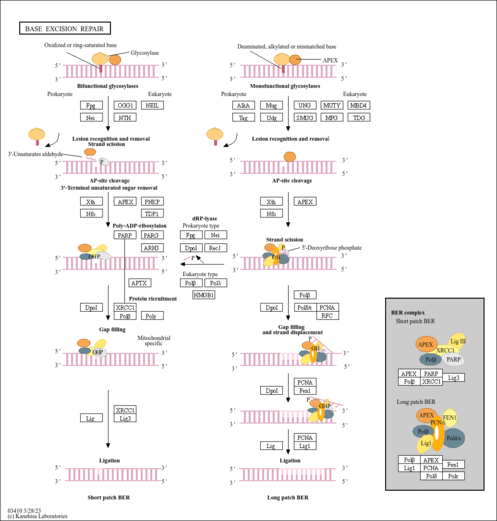 | | NF-kappa B signaling pathway | Not Available | 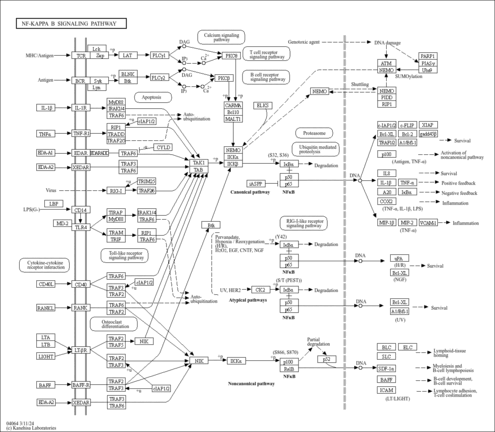 |
|
| Gene Properties |
|---|
| Chromosome Location
| 1 |
| Locus
| 1q41-q42 |
| SNPs
| PARP1 |
| Gene Sequence
|
>3045 bp
ATGGCGGAGTCTTCGGATAAGCTCTATCGAGTCGAGTACGCCAAGAGCGGGCGCGCCTCT
TGCAAGAAATGCAGCGAGAGCATCCCCAAGGACTCGCTCCGGATGGCCATCATGGTGCAG
TCGCCCATGTTTGATGGAAAAGTCCCACACTGGTACCACTTCTCCTGCTTCTGGAAGGTG
GGCCACTCCATCCGGCACCCTGACGTTGAGGTGGATGGGTTCTCTGAGCTTCGGTGGGAT
GACCAGCAGAAAGTCAAGAAGACAGCGGAAGCTGGAGGAGTGACAGGCAAAGGCCAGGAT
GGAATTGGTAGCAAGGCAGAGAAGACTCTGGGTGACTTTGCAGCAGAGTATGCCAAGTCC
AACAGAAGTACGTGCAAGGGGTGTATGGAGAAGATAGAAAAGGGCCAGGTGCGCCTGTCC
AAGAAGATGGTGGACCCGGAGAAGCCACAGCTAGGCATGATTGACCGCTGGTACCATCCA
GGCTGCTTTGTCAAGAACAGGGAGGAGCTGGGTTTCCGGCCCGAGTACAGTGCGAGTCAG
CTCAAGGGCTTCAGCCTCCTTGCTACAGAGGATAAAGAAGCCCTGAAGAAGCAGCTCCCA
GGAGTCAAGAGTGAAGGAAAGAGAAAAGGCGATGAGGTGGATGGAGTGGATGAAGTGGCG
AAGAAGAAATCTAAAAAAGAAAAAGACAAGGATAGTAAGCTTGAAAAAGCCCTAAAGGCT
CAGAACGACCTGATCTGGAACATCAAGGACGAGCTAAAGAAAGTGTGTTCAACTAATGAC
CTGAAGGAGCTACTCATCTTCAACAAGCAGCAAGTGCCTTCTGGGGAGTCGGCGATCTTG
GACCGAGTAGCTGATGGCATGGTGTTCGGTGCCCTCCTTCCCTGCGAGGAATGCTCGGGT
CAGCTGGTCTTCAAGAGCGATGCCTATTACTGCACTGGGGACGTCACTGCCTGGACCAAG
TGTATGGTCAAGACACAGACACCCAACCGGAAGGAGTGGGTAACCCCAAAGGAATTCCGA
GAAATCTCTTACCTCAAGAAATTGAAGGTTAAAAAGCAGGACCGTATATTCCCCCCAGAA
ACCAGCGCCTCCGTGGCGGCCACGCCTCCGCCCTCCACAGCCTCGGCTCCTGCTGCTGTG
AACTCCTCTGCTTCAGCAGATAAGCCATTATCCAACATGAAGATCCTGACTCTCGGGAAG
CTGTCCCGGAACAAGGATGAAGTGAAGGCCATGATTGAGAAACTCGGGGGGAAGTTGACG
GGGACGGCCAACAAGGCTTCCCTGTGCATCAGCACCAAAAAGGAGGTGGAAAAGATGAAT
AAGAAGATGGAGGAAGTAAAGGAAGCCAACATCCGAGTTGTGTCTGAGGACTTCCTCCAG
GACGTCTCCGCCTCCACCAAGAGCCTTCAGGAGTTGTTCTTAGCGCACATCTTGTCCCCT
TGGGGGGCAGAGGTGAAGGCAGAGCCTGTTGAAGTTGTGGCCCCAAGAGGGAAGTCAGGG
GCTGCGCTCTCCAAAAAAAGCAAGGGCCAGGTCAAGGAGGAAGGTATCAACAAATCTGAA
AAGAGAATGAAATTAACTCTTAAAGGAGGAGCAGCTGTGGATCCTGATTCTGGACTGGAA
CACTCTGCGCATGTCCTGGAGAAAGGTGGGAAGGTCTTCAGTGCCACCCTTGGCCTGGTG
GACATCGTTAAAGGAACCAACTCCTACTACAAGCTGCAGCTTCTGGAGGACGACAAGGAA
AACAGGTATTGGATATTCAGGTCCTGGGGCCGTGTGGGTACGGTGATCGGTAGCAACAAA
CTGGAACAGATGCCGTCCAAGGAGGATGCCATTGAGCACTTCATGAAATTATATGAAGAA
AAAACCGGGAACGCTTGGCACTCCAAAAATTTCACGAAGTATCCCAAAAAGTTCTACCCC
CTGGAGATTGACTATGGCCAGGATGAAGAGGCAGTGAAGAAGCTGACAGTAAATCCTGGC
ACCAAGTCCAAGCTCCCCAAGCCAGTTCAGGACCTCATCAAGATGATCTTTGATGTGGAA
AGTATGAAGAAAGCCATGGTGGAGTATGAGATCGACCTTCAGAAGATGCCCTTGGGGAAG
CTGAGCAAAAGGCAGATCCAGGCCGCATACTCCATCCTCAGTGAGGTCCAGCAGGCGGTG
TCTCAGGGCAGCAGCGACTCTCAGATCCTGGATCTCTCAAATCGCTTTTACACCCTGATC
CCCCACGACTTTGGGATGAAGAAGCCTCCGCTCCTGAACAATGCAGACAGTGTGCAGGCC
AAGGTGGAAATGCTTGACAACCTGCTGGACATCGAGGTGGCCTACAGTCTGCTCAGGGGA
GGGTCTGATGATAGCAGCAAGGATCCCATCGATGTCAACTATGAGAAGCTCAAAACTGAC
ATTAAGGTGGTTGACAGAGATTCTGAAGAAGCCGAGATCATCAGGAAGTATGTTAAGAAC
ACTCATGCAACCACACACAATGCGTATGACTTGGAAGTCATCGATATCTTTAAGATAGAG
CGTGAAGGCGAATGCCAGCGTTACAAGCCCTTTAAGCAGCTTCATAACCGAAGATTGCTG
TGGCACGGGTCCAGGACCACCAACTTTGCTGGGATCCTGTCCCAGGGTCTTCGGATAGCC
CCGCCTGAAGCGCCCGTGACAGGCTACATGTTTGGTAAAGGGATCTATTTCGCTGACATG
GTCTCCAAGAGTGCCAACTACTGCCATACGTCTCAGGGAGACCCAATAGGCTTAATCCTG
TTGGGAGAAGTTGCCCTTGGAAACATGTATGAACTGAAGCACGCTTCACATATCAGCAAG
TTACCCAAGGGCAAGCACAGTGTCAAAGGTTTGGGCAAAACTACCCCTGATCCTTCAGCT
AACATTAGTCTGGATGGTGTAGACGTTCCTCTTGGGACCGGGATTTCATCTGGTGTGAAT
GACACCTCTCTACTATATAACGAGTACATTGTCTATGATATTGCTCAGGTAAATCTGAAG
TATCTGCTGAAACTGAAATTCAATTTTAAGACCTCCCTGTGGTAA
|
| Protein Properties |
|---|
| Number of Residues
| 1014 |
| Molecular Weight
| 113082.945 |
| Theoretical pI
| 8.885 |
| Pfam Domain Function
|
|
| Signals
|
Not Available
|
|
Transmembrane Regions
|
Not Available
|
| Protein Sequence
|
>Poly [ADP-ribose] polymerase 1
MAESSDKLYRVEYAKSGRASCKKCSESIPKDSLRMAIMVQSPMFDGKVPHWYHFSCFWKV
GHSIRHPDVEVDGFSELRWDDQQKVKKTAEAGGVTGKGQDGIGSKAEKTLGDFAAEYAKS
NRSTCKGCMEKIEKGQVRLSKKMVDPEKPQLGMIDRWYHPGCFVKNREELGFRPEYSASQ
LKGFSLLATEDKEALKKQLPGVKSEGKRKGDEVDGVDEVAKKKSKKEKDKDSKLEKALKA
QNDLIWNIKDELKKVCSTNDLKELLIFNKQQVPSGESAILDRVADGMVFGALLPCEECSG
QLVFKSDAYYCTGDVTAWTKCMVKTQTPNRKEWVTPKEFREISYLKKLKVKKQDRIFPPE
TSASVAATPPPSTASAPAAVNSSASADKPLSNMKILTLGKLSRNKDEVKAMIEKLGGKLT
GTANKASLCISTKKEVEKMNKKMEEVKEANIRVVSEDFLQDVSASTKSLQELFLAHILSP
WGAEVKAEPVEVVAPRGKSGAALSKKSKGQVKEEGINKSEKRMKLTLKGGAAVDPDSGLE
HSAHVLEKGGKVFSATLGLVDIVKGTNSYYKLQLLEDDKENRYWIFRSWGRVGTVIGSNK
LEQMPSKEDAIEHFMKLYEEKTGNAWHSKNFTKYPKKFYPLEIDYGQDEEAVKKLTVNPG
TKSKLPKPVQDLIKMIFDVESMKKAMVEYEIDLQKMPLGKLSKRQIQAAYSILSEVQQAV
SQGSSDSQILDLSNRFYTLIPHDFGMKKPPLLNNADSVQAKVEMLDNLLDIEVAYSLLRG
GSDDSSKDPIDVNYEKLKTDIKVVDRDSEEAEIIRKYVKNTHATTHNAYDLEVIDIFKIE
REGECQRYKPFKQLHNRRLLWHGSRTTNFAGILSQGLRIAPPEAPVTGYMFGKGIYFADM
VSKSANYCHTSQGDPIGLILLGEVALGNMYELKHASHISKLPKGKHSVKGLGKTTPDPSA
NISLDGVDVPLGTGISSGVNDTSLLYNEYIVYDIAQVNLKYLLKLKFNFKTSLW
|
| External Links |
|---|
| GenBank ID Protein
| 21693601 |
| UniProtKB/Swiss-Prot ID
| P09874 |
| UniProtKB/Swiss-Prot Entry Name
| PARP1_HUMAN |
| PDB IDs
|
|
| GenBank Gene ID
| AF524947 |
| GeneCard ID
| PARP1 |
| GenAtlas ID
| PARP1 |
| HGNC ID
| HGNC:270 |
| References |
|---|
| General References
| Not Available |

