| Identification |
|---|
| HMDB Protein ID
| CDBP00815 |
| Secondary Accession Numbers
| Not Available |
| Name
| Beta-hexosaminidase subunit alpha |
| Description
| Not Available |
| Synonyms
|
- Beta-N-acetylhexosaminidase subunit alpha
- Hexosaminidase subunit A
- N-acetyl-beta-glucosaminidase subunit alpha
|
| Gene Name
| HEXA |
| Protein Type
| Enzyme |
| Biological Properties |
|---|
| General Function
| Involved in beta-N-acetylhexosaminidase activity |
| Specific Function
| Responsible for the degradation of GM2 gangliosides, and a variety of other molecules containing terminal N-acetyl hexosamines, in the brain and other tissues. The form B is active against certain oligosaccharides. The form S has no measurable activity.
|
| GO Classification
|
| Biological Process |
| adult walking behavior |
| neuromuscular process controlling posture |
| skeletal system development |
| cell morphogenesis involved in neuron differentiation |
| ganglioside catabolic process |
| sexual reproduction |
| carbohydrate metabolic process |
| glycosphingolipid metabolic process |
| cell death |
| myelination |
| chondroitin sulfate catabolic process |
| sensory perception of sound |
| lysosome organization |
| neuromuscular process controlling balance |
| lipid storage |
| phospholipid metabolic process |
| keratan sulfate catabolic process |
| hyaluronan catabolic process |
| Cellular Component |
| lysosomal lumen |
| membrane |
| Function |
| beta-n-acetylhexosaminidase activity |
| binding |
| catalytic activity |
| hydrolase activity |
| hydrolase activity, acting on glycosyl bonds |
| hydrolase activity, hydrolyzing o-glycosyl compounds |
| ion binding |
| cation binding |
| hexosaminidase activity |
| Molecular Function |
| cation binding |
| beta-N-acetylhexosaminidase activity |
| protein heterodimerization activity |
| Process |
| metabolic process |
| primary metabolic process |
| carbohydrate metabolic process |
|
| Cellular Location
|
- Lysosome
|
| Pathways
|
| Name | SMPDB/Pathwhiz | KEGG | | Other glycan degradation | Not Available | 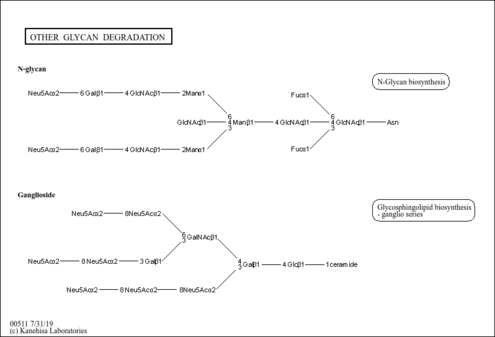 | | Amino sugar and nucleotide sugar metabolism | Not Available | 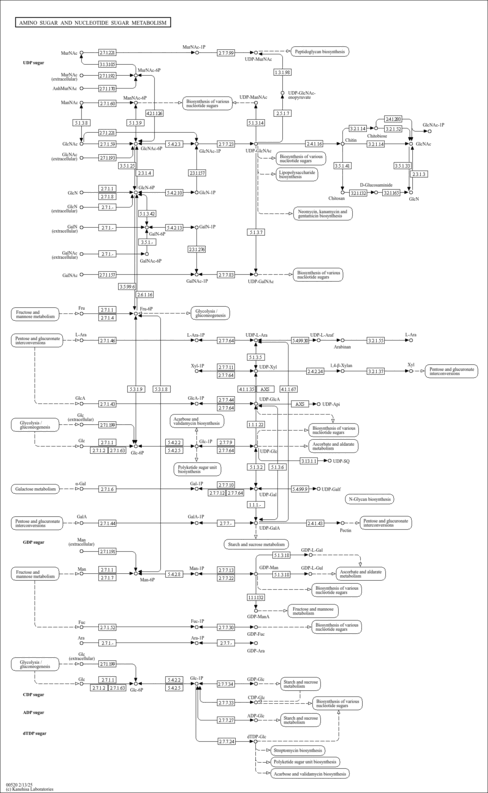 | | Glycosaminoglycan degradation | Not Available | 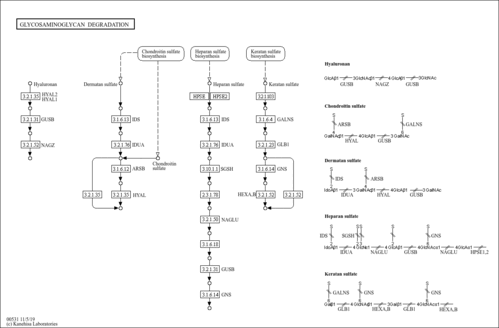 | | Glycosphingolipid biosynthesis - globo series | Not Available | 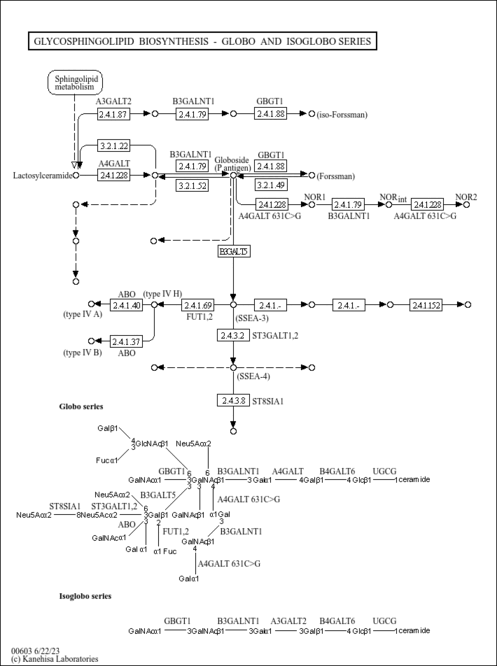 | | Glycosphingolipid biosynthesis - ganglio series | Not Available | 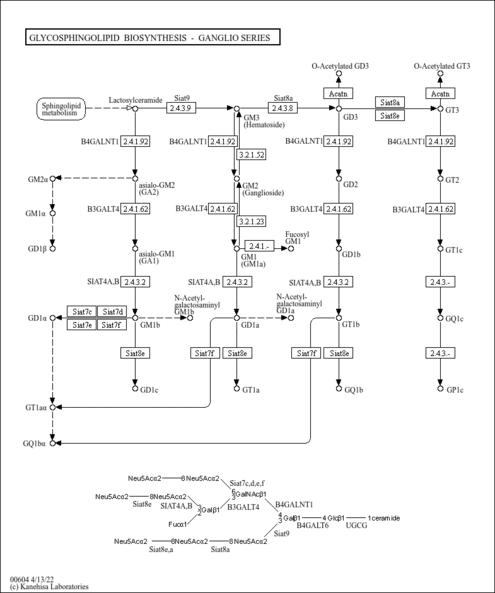 |
|
| Gene Properties |
|---|
| Chromosome Location
| Not Available |
| Locus
| Not Available |
| SNPs
| HEXA |
| Gene Sequence
|
>1590 bp
ATGACAAGCTCCAGGCTTTGGTTTTCGCTGCTGCTGGCGGCAGCGTTCGCAGGACGGGCG
ACGGCCCTCTGGCCCTGGCCTCAGAACTTCCAAACCTCCGACCAGCGCTACGTCCTTTAC
CCGAACAACTTTCAATTCCAGTACGATGTCAGCTCGGCCGCGCAGCCCGGCTGCTCAGTC
CTCGACGAGGCCTTCCAGCGCTATCGTGACCTGCTTTTCGGTTCCGGGTCTTGGCCCCGT
CCTTACCTCACAGGGAAACGGCATACACTGGAGAAGAATGTGTTGGTTGTCTCTGTAGTC
ACACCTGGATGTAACCAGCTTCCTACTTTGGAGTCAGTGGAGAATTATACCCTGACCATA
AATGATGACCAGTGTTTACTCCTCTCTGAGACTGTCTGGGGAGCTCTCCGAGGTCTGGAG
ACTTTTAGCCAGCTTGTTTGGAAATCTGCTGAGGGCACATTCTTTATCAACAAGACTGAG
ATTGAGGACTTTCCCCGCTTTCCTCACCGGGGCTTGCTGTTGGATACATCTCGCCATTAC
CTGCCACTCTCTAGCATCCTGGACACTCTGGATGTCATGGCGTACAATAAATTGAACGTG
TTCCACTGGCATCTGGTAGATGATCCTTCCTTCCCATATGAGAGCTTCACTTTTCCAGAG
CTCATGAGAAAGGGGTCCTACAACCCTGTCACCCACATCTACACAGCACAGGATGTGAAG
GAGGTCATTGAATACGCACGGCTCCGGGGTATCCGTGTGCTTGCAGAGTTTGACACTCCT
GGCCACACTTTGTCCTGGGGACCAGGTATCCCTGGATTACTGACTCCTTGCTACTCTGGG
TCTGAGCCCTCTGGCACCTTTGGACCAGTGAATCCCAGTCTCAATAATACCTATGAGTTC
ATGAGCACATTCTTCTTAGAAGTCAGCTCTGTCTTCCCAGATTTTTATCTTCATCTTGGA
GGAGATGAGGTTGATTTCACCTGCTGGAAGCCCAACCCAGAGATCCAGGACTTTATGAGG
AAGAAAGGCTTCGGTGAGGACTTCAAGCAGCTGGAGTCCTTCTACATCCAGACGCTGCTG
GACATCGTCTCTTCTTATGGCAAGGGCTATGTGGTGTGGCAGGAGGTGTTTGATAATAAA
GTAAAGATTCAGCCAGACACAATCATACAGGTGTGGCGAGAGGATATTCCAGTGAACTAT
ATGAAGGAGCTGGAACTGGTCACCAAGGCCGGCTTCCGGGCCCTTCTCTCTGCCCCCTGG
TACCTGAACCGTATATCCTACGGCCCTGACTGGAAGGATTTCTACGTAGTGGAACCCCTG
GCATTTGAAGGTACCCCTGAGCAGAAGGCTCTGGTGATTGGTGGAGAGGCTTGTATGTGG
GGAGAATATGTGGACAACACAAACCTGGTCCCCAGGCTCTGGCCCAGAGCAGGGGCTGTT
GCCGAAAGGCTGTGGAGCAACAAGTTGACATCTGACCTGACATTTGCCTATGAACGTTTG
TCACACTTCCGCTGTGAGTTGCTGAGGCGAGGTGTCCAGGCCCAACCCCTCAATGTAGGC
TTCTGTGAGCAGGAGTTTGAACAGACCTGA
|
| Protein Properties |
|---|
| Number of Residues
| 529 |
| Molecular Weight
| Not Available |
| Theoretical pI
| Not Available |
| Pfam Domain Function
|
|
| Signals
|
Not Available
|
|
Transmembrane Regions
|
Not Available
|
| Protein Sequence
|
>Beta-hexosaminidase subunit alpha
MTSSRLWFSLLLAAAFAGRATALWPWPQNFQTSDQRYVLYPNNFQFQYDVSSAAQPGCSV
LDEAFQRYRDLLFGSGSWPRPYLTGKRHTLEKNVLVVSVVTPGCNQLPTLESVENYTLTI
NDDQCLLLSETVWGALRGLETFSQLVWKSAEGTFFINKTEIEDFPRFPHRGLLLDTSRHY
LPLSSILDTLDVMAYNKLNVFHWHLVDDPSFPYESFTFPELMRKGSYNPVTHIYTAQDVK
EVIEYARLRGIRVLAEFDTPGHTLSWGPGIPGLLTPCYSGSEPSGTFGPVNPSLNNTYEF
MSTFFLEVSSVFPDFYLHLGGDEVDFTCWKSNPEIQDFMRKKGFGEDFKQLESFYIQTLL
DIVSSYGKGYVVWQEVFDNKVKIQPDTIIQVWREDIPVNYMKELELVTKAGFRALLSAPW
YLNRISYGPDWKDFYVVEPLAFEGTPEQKALVIGGEACMWGEYVDNTNLVPRLWPRAGAV
AERLWSNKLTSDLTFAYERLSHFRCELLRRGVQAQPLNVGFCEQEFEQT
|
| External Links |
|---|
| GenBank ID Protein
| 62896563 |
| UniProtKB/Swiss-Prot ID
| P06865 |
| UniProtKB/Swiss-Prot Entry Name
| HEXA_HUMAN |
| PDB IDs
|
|
| GenBank Gene ID
| AK222502 |
| GeneCard ID
| HEXA |
| GenAtlas ID
| HEXA |
| HGNC ID
| HGNC:4878 |
| References |
|---|
| General References
| Not Available |




